R&D Success: Revolutionary genome sequencing of soy sauce koji mold


Soy sauce: The miracle seasoning made by microorganisms
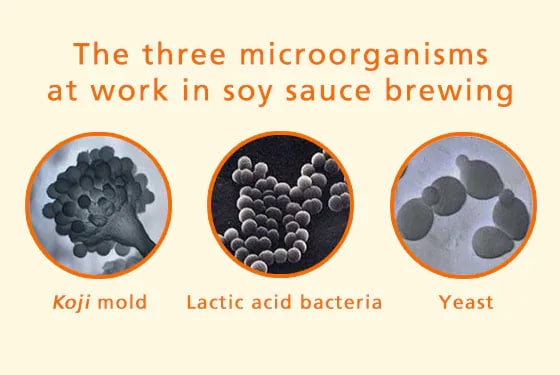
Soybeans, wheat, and salt are the raw ingredients used to produce soy sauce. At the same time, within the soy sauce brewing process, three kinds of microorganism are also involved. These are: koji molds, lactic acid bacteria, and yeasts. Soy sauce can be thought of as a kind of miracle seasoning which can be manufactured thanks to the action of these three types of microorganism.
Obtaining the DNA draft genome sequences of the essential microorganisms!
We skillfully harness the action of these three kinds of microorganisms to manufacture Kikkoman Soy Sauce. However, there remain aspects of the action of living microorganisms which have yet to be scientifically clarified, meaning that there are elements of this process which rely on our experience alone.
All living organisms possess genetic information called the genome, which is often described as “blueprint” for life. We embarked on the challenge to sequence the genome of the microorganisms involved in soy sauce brewing with the reasoning that by sequencing (1) and understanding the characteristics of these genomes we might gain a scientific grasp of the particular means of action of these microorganisms.

Tell me ALL ABOUT “genomes” (1)? Just WHAT is “sequencing”?
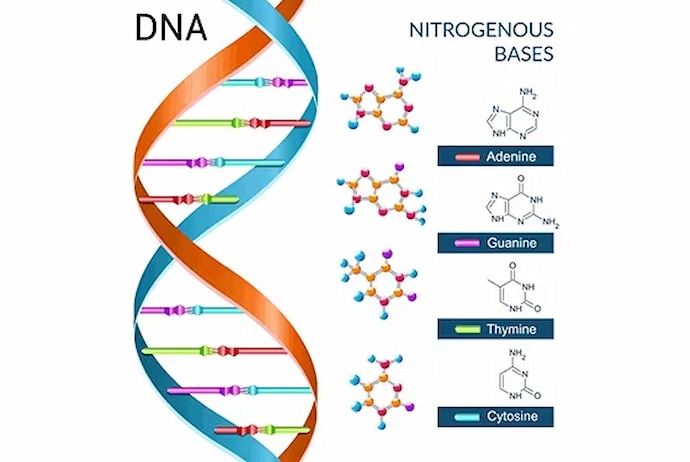
A genome is comprised of only four substances (nucleotides): “adenine”, “thymine”, “guanine” and “cytosine”. Genetic information is determined by the arrangement of these nucleotides. Whole genome sequencing uses an instrument called a sequencer to determine the sequence in which these nucleotides are arranged.
Our story began with decoding the sequence of Aspergillus oryzae
There are two kinds of koji mold used to make the essential base of soy sauce: Aspergillus oryzae and A. sojae. Kikkoman participated in the genome sequencing project of A. oryzae, an undertaking which was conducted in conjunction with a variety of private sector companies, involved with the soy sauce, sake and enzyme industries, as well as universities and research institutes.
Subsequently, the sequencing of A. sojae was conducted based on the genetic information obtained from the sequencing of A. oryzae. However, it became increasingly apparent as this process evolved that the two were considerably more at variance than was initially assumed. Thus, further research was required.
Genome sequencing of A. sojae

We cooperated with the Noda Institute for Scientific Research, University of Tokyo, and Tokyo Institute of Technology to read the A. sojae genome using a next-generation genome sequencing (2) technology. Six months after embarking on this endeavor we had successfully sequenced the whole genome and presented the outcomes of our efforts in a scientific paper.
By sequencing the genome, we were able to confirm that A. oryzae and A. sojae is not capable of producing mycotoxins at gene level, and thus obtained the evidence supporting the safety for human consumption.

Just WHAT IS a next-generation genome sequencer?
A next-generation genome sequencer is a new type of device designed to elucidate DNA sequence both speedily and at low cost. It achieves speeds of several thousand times those of conventional sequencers.
Furthering the scientific understanding of soy sauce brewing

There is little to be gained by the mere act of sequencing a genome. Genome information is of vast scales similar to big data, and bioinformatics technologies are needed to mine the useful information for particular purposes. At present, we use bioinformatics to analyze the genome information of the three kinds of microorganisms involved in the brewing process for soy sauce, to systematically elucidate the properties of these microorganisms and their role in soy sauce brewing. Our wish is to uncover the science behind soy sauce brewing to improve our products to be loved even more by people throughout the globe.
This research was awarded the Soy Sauce Technology Award (R&D Section) of the Japan Soy Sauce Brewers’ Association / Japan Soy Sauce Technology Center in Japan.
Coverage in external publications
Sato, A., Oshima, K., Noguchi, H., Ogawa, M., Takahashi, T., Oguma, T., Koyama, Y., Itoh, T., Hattori, M., & Hanya, Y. (2011). Draft genome sequencing and comparative analysis of Aspergillus sojae NBRC4239. DNA research, 18(3), 165-176.
